Estimates the noise standard deviation (\(\sigma\)) for each spectrum from a signal-free ppm region. The default estimator is robust (MAD-based) and suitable for signal-to-noise calculations.
Arguments
- X
Numeric matrix (spectra in rows), numeric vector (single spectrum), or a named list as returned by
read1d/read1d_proccontainingX,ppm, andmeta.- ppm
Numeric vector of chemical shift values (ppm) corresponding to columns of
X. IfNULL,ppmis inferred in the following order:X$ppmifXis a list input,attr(X, "m8_axis")$ppm(if present),numeric
colnames(X)(if present).
- where
Numeric vector of length 2. ppm range used for noise estimation. Should be free of metabolite signals (default
c(14.6, 14.7)).- method
Character. Noise estimator:
"mad"(default),"sd", or"p95"(legacy amplitude).- baseline_correct
Logical. If
TRUE, subtract a smooth baseline in the noise window using asymmetric least squares (asysm). DefaultFALSE.- lambda
Numeric. Smoothing parameter for
asysmwhenbaseline_correct = TRUE.- min_points
Integer. Minimum number of points required in the noise window. May be a single value (recycled across spectra) or a numeric vector of length
nrow(X). IfXis a list input as returned byread1d/read1d_procandnsisNULL, the function attempts to extract the number of scans fromX$meta$a_NSwhen available.- ns
Number of scans / transients (required if normalise_scans=TRUE)
- normalise_scans
Logical. If
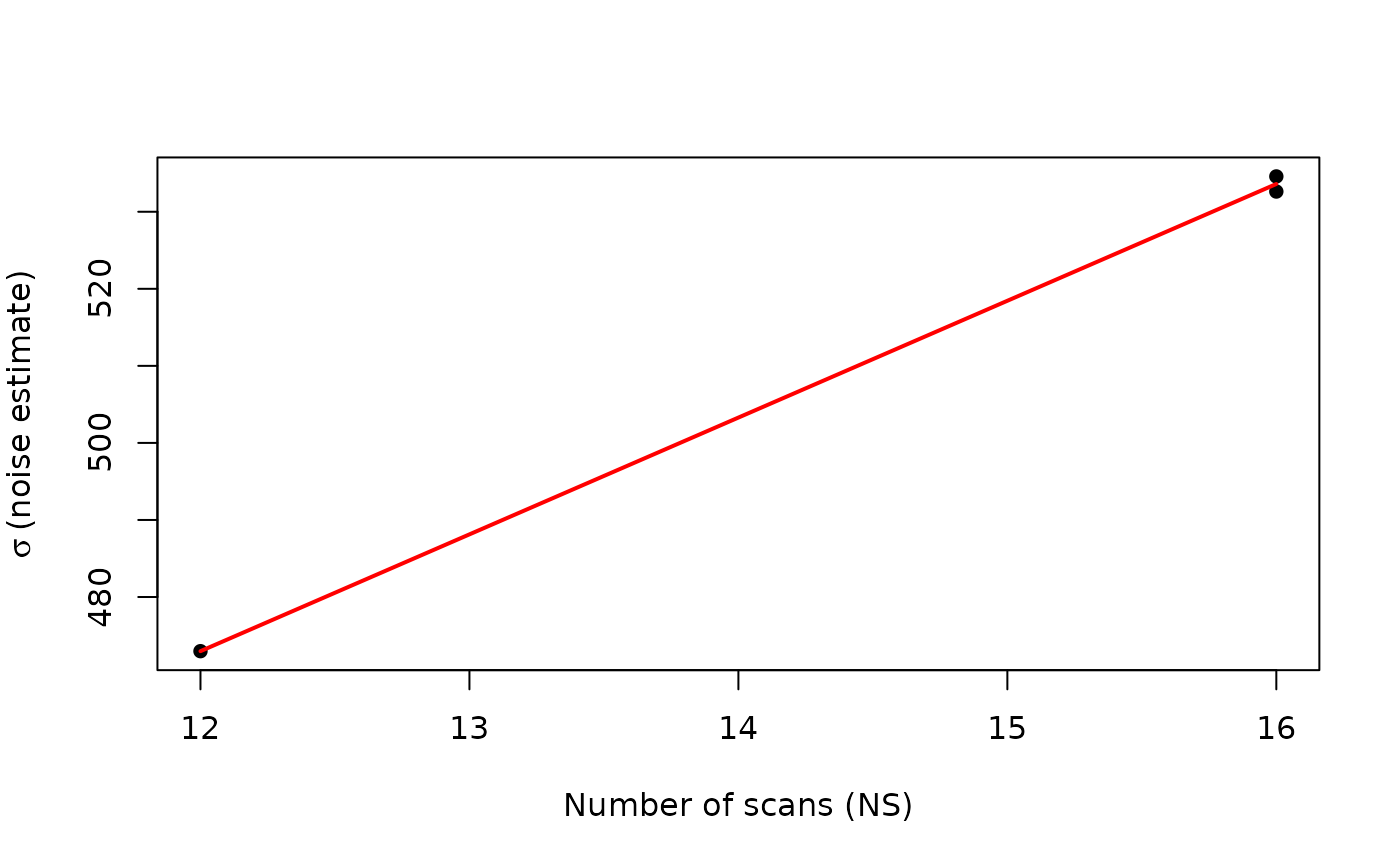
TRUE, noise estimates are multiplied by \(\sqrt{NS}\) to account for the theoretical scaling of noise with the number of scans (\(\sigma \propto 1/\sqrt{NS}\)). This is useful when comparing noise levels across spectra acquired with different numbers of scans. Default isFALSE.
Details
In NMR spectroscopy, noise scales predictably with the number of scans (\(NS\)). For otherwise identical acquisition settings:
Signal increases approximately proportional to \(NS\).
Noise increases approximately proportional to \(\sqrt{NS}\).
Consequently, signal-to-noise ratio (SNR) increases proportional to \(\sqrt{NS}\).
Equivalently, the noise standard deviation scales as:
$$\sigma(NS) \propto \frac{1}{\sqrt{NS}},$$
assuming a fixed underlying signal scale and comparable acquisition conditions.
To compare noise levels across datasets acquired with different numbers of scans, a scan-normalised noise estimate may be used:
$$\sigma_{\mathrm{norm}} = \sigma \cdot \sqrt{NS}.$$
Under stable receiver gain and processing conditions, this normalised noise should be approximately constant across runs.